A few words of caution
You should be careful when interpreting the returned predictions for several reasons. In the following I will explain some of the pitfalls of interpreting PredRet predictions.
Contents
Prediction intervals
Determining the prediction intervals
When retention time (RT) predictions are used to annotate untargeted data sets, it is useful to know not just the predicted RT but also a prediction interval (PI). PIs are unfortunately not easily established for all but the simplest models, and for this reason we found that of the previously developed RT prediction methods only the models based on a linear relationship between log P and RT provide PIs and not for individual predictions as such.35 Likewise, we are currently not aware of methods for building PIs for GAM models with constraints. We therefore used bootstrapping to establish empirical PIs.
When the PIs for each prediction are joined we get prediction bands. Correctly defined prediction bands would ensure that the fraction of predictions where the true RTs fall outside the prediction band is no more than the chosen confidence level. In our case, however, because of multiplicity issues this cannot be ensured. PIs should therefore be regarded as anticonservative and not be used as strict filters to exclude a possible compound annotation but rather as an indication of the likelihood of a match. The accuracy of the predictions and width of the PIs depend on the accuracy of the projection model associated with each prediction.
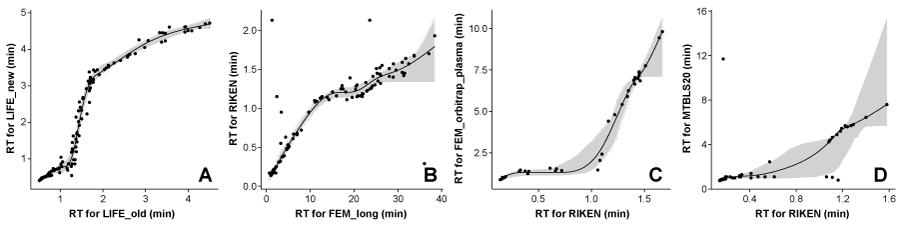
In Figure 1A the model used to predict RTs from the CS “LIFE_old” to the CS “LIFE_new” can be seen. This model is very accurate since there are no major outliers and many compounds with known RT in both CSs. This leads to very narrow PIs for the predictions. It should be noted, however, that due to the lack of methods to construct accurate prediction bands as explained above, the very narrow PIs can be misleading. In Figure 1A, for the interval around 1.5 min (for LIFE_old) the PI is narrow in part because the slope is steep, which makes it easier to establish the fitted curve. Unfortunately a steep slope also means that relatively small inaccuracies (instrumental variation) in the independent variable (LIFE_old RT) leads to relatively large errors in the prediction of the dependent variable (LIFE_new RT) which the PI does not reflect.
We will thus reiterate that the PIs are anticonservative and should be approached as such. Nevertheless, we believe that providing anticonservative PIs are better than having no PIs at all.
Changes over time
Another potential source of inaccuracies, that all prediction systems suffer from, is the potential for small changes in RTs over time. This can be caused by column aging but also by small changes in the eluent composition such as slight pH differences. These changes are not necessarily systematic and can therefore increase the width of the PIs if the set of RTs used for modeling was accumulated over time. In addition the accuracy of the predictions might be lower than expected based on the model if the “current” system differs sufficiently from the system the original RTs were recorded on.
Errors in the database entries
The predicted retention time depends on reported retention times from other people. People make mistakes both of the copy/paste type but also in correct identification. While we do try to detect erroneous there is no guarantee that predictions are not made from erroneous reports that can cause a predicted retention time to be far off.
Unambiguous molecular structures
Some compounds are often reported without defining an unambiguous molecular structure. Lets take a common example like LysoPC(18:1). There are at least three structural aspect that are not defined by the name and often unknown when analyzing untargeted data or even targeted data:
- The lipid chain can be in two different positions on the glycerol, sn1 (i.e., LysoPC(18:1/0:0)) and sn2 (i.e., LysoPC(0:0/18:1)).
- The position of the double bond is not specified.
- The relative stereochemistry around the double bond is not specified (i.e., cis/trans).
Differences in any of these aspects can lead to different RTs.
This is not a limitation of PredRet as such but a limitation of current reporting standards.
Diastereoisomers
Since stereochemistry in general cannot be determined in LC, it is ignored by PredRet. In general this has no influence on the predicted RT since enantiomers have the same RT on nonchiral columns.
Diastereoisomers, however, do not necessarily have the same RT. An example is hydroxy-methyl- butyric acid for which there exist a number of structural isomers (PubChem CIDs: 95433, 99823, 160471, 69362, 14081034, 131760, 188979). Most of them have at least one chiral center and therefore exists in enantiomeric forms.
3-Hydroxy-2-methylbutyric acid on the other hand has two chiral centers and thus exists in four forms that are enantiomers and diastereoisomers in pairs of two (PubChem CIDs: 12313369, 11966260 and 11815846, 12313370). The diastereoisomers can have different RTs while the enantiomers cannot. The PI of a prediction for 3-hydroxy-2-methylbutyric acid might therefore not match the experimental RT if the database entry and the experimental data was obtained from different diastereoisomers.
